As our understanding of fish larvae dispersal improves, biophysical models become more and more complex - a necessary step to accurately reproduce the behaviours displayed by fish larvae during their pelagic life. The utility of these biophysical models in informing decision-makers for the establishment of marine protected areas (MPAs) or advising sustainable fisheries management depends on the accuracy of the models' estimates. These complex models, unfortunately, rely on input parameters and equations that are poorly understood and for which quantitative studies are lacking. More studies are needed to quantify the sensitivity of the models to uncertain parameters and assess their overall reliability.
The latest Editor's Choice article in ICES Journal of Marine Science presents a novel application of the Polynomial Chaos method to quantify uncertainties. The method helped in estimating the impact of biological uncertainties associated with a complex biophysical model of fish larvae (Abudefduf saxatilis) dispersal in the Florida Keys. The authors of the current study used results of recent publications to improve a model (i.e. Connectivity Modeling System) with a module that simulates fish larval orientation behaviour. This behaviour has been shown to modify connectivity estimates, but its implementation in biophysical models requires multiple biological parameters - some well-described in the literature, while others are seldomly studied.
Instead of arbitrability fixing values for these parameters, Chaput et al. considered their observed or supposed distributions to investigate their impact on the model outputs. Multiple simulations were run to create a Polynomial Chaos surrogate that directly relates the biological parameters to the uncertainty in connectivity estimates. A global sensitivity analysis showed that the interaction between a few parameters - associated with the orientation behavior - are the dominant contributors to the uncertainty in the modeled larvae dispersal. On the other hand, parameters that have been the focus of previous research and experimentation, such as swimming speed, seem to contribute little to dispersal uncertainties.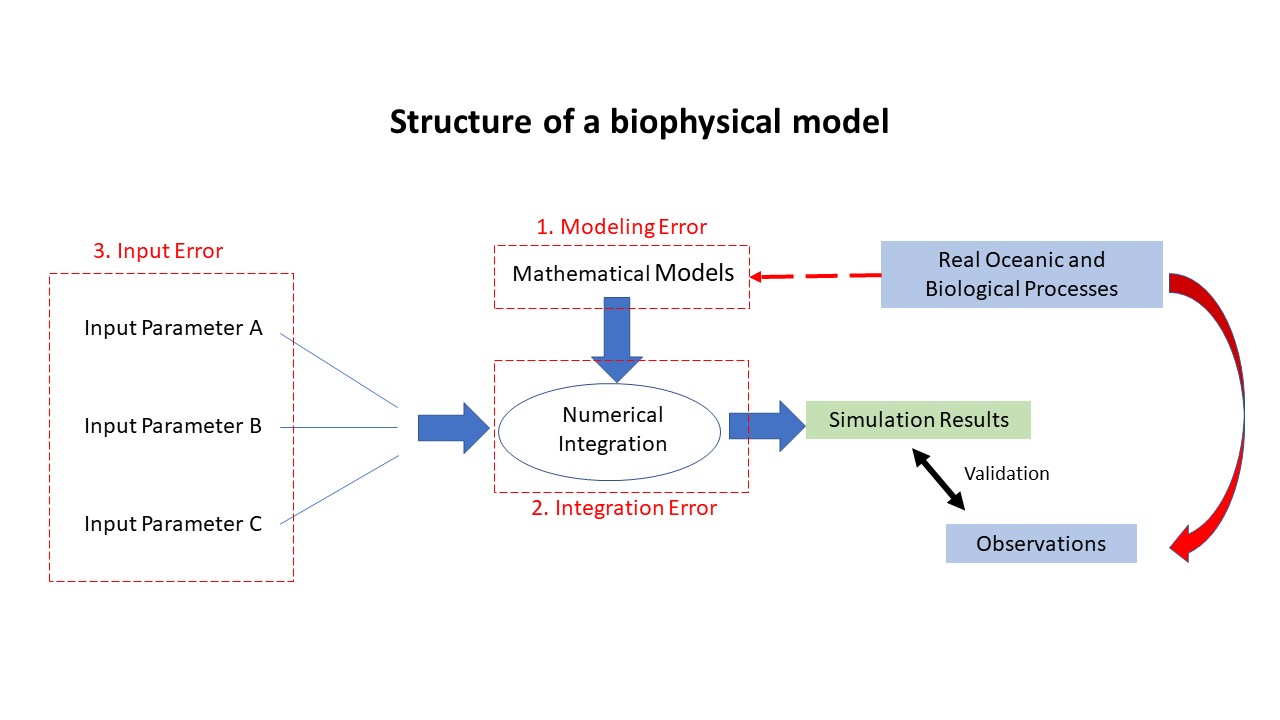
Structure of a biophysical model and the three categories of associated errors. Uncertainties in forecast model, such as biophysical models of larval fish, can be categorized into three classes: 1. model uncertainties caused by the abstraction of the true processes (physical or biological) into mathematical equations; 2. uncertainties due to the numerical integration; and 3. input data uncertainties due to the empirical parameters involved in the model itself. The focus of this study is the characterization of the third type of uncertainties.
The Polynomial Chaos method adopted in the study to quantify the uncertainties increases the reliability of the output and provides the user with an estimate of the model usefulness, while keeping the computational cost to a minimum when compared to other quantification methods. Finally, this research highlights the need to study the orientation behaviors of fish larvae with a quantitative approach rather than the descriptive approach used to date.
Read the full paper, Quantitative uncertainty estimation in biophysical models of fish larval connectivity in the Florida Keys, in ICES Journal of Marine Science.
Editor's Choice articles are always free to read in ICES Journal of Marine Science .

